Sample preparation is the #1 bottleneck in high-throughput proteomics. It's also the #1 source of error.
Your mass spec can process hundreds of samples a day. Your sample prep can't match that pace. PIXUL changes that. 96 samples in parallel, in 10–30 minutes, with LC-MS/MS-validated protein identification equivalent to probe sonication.
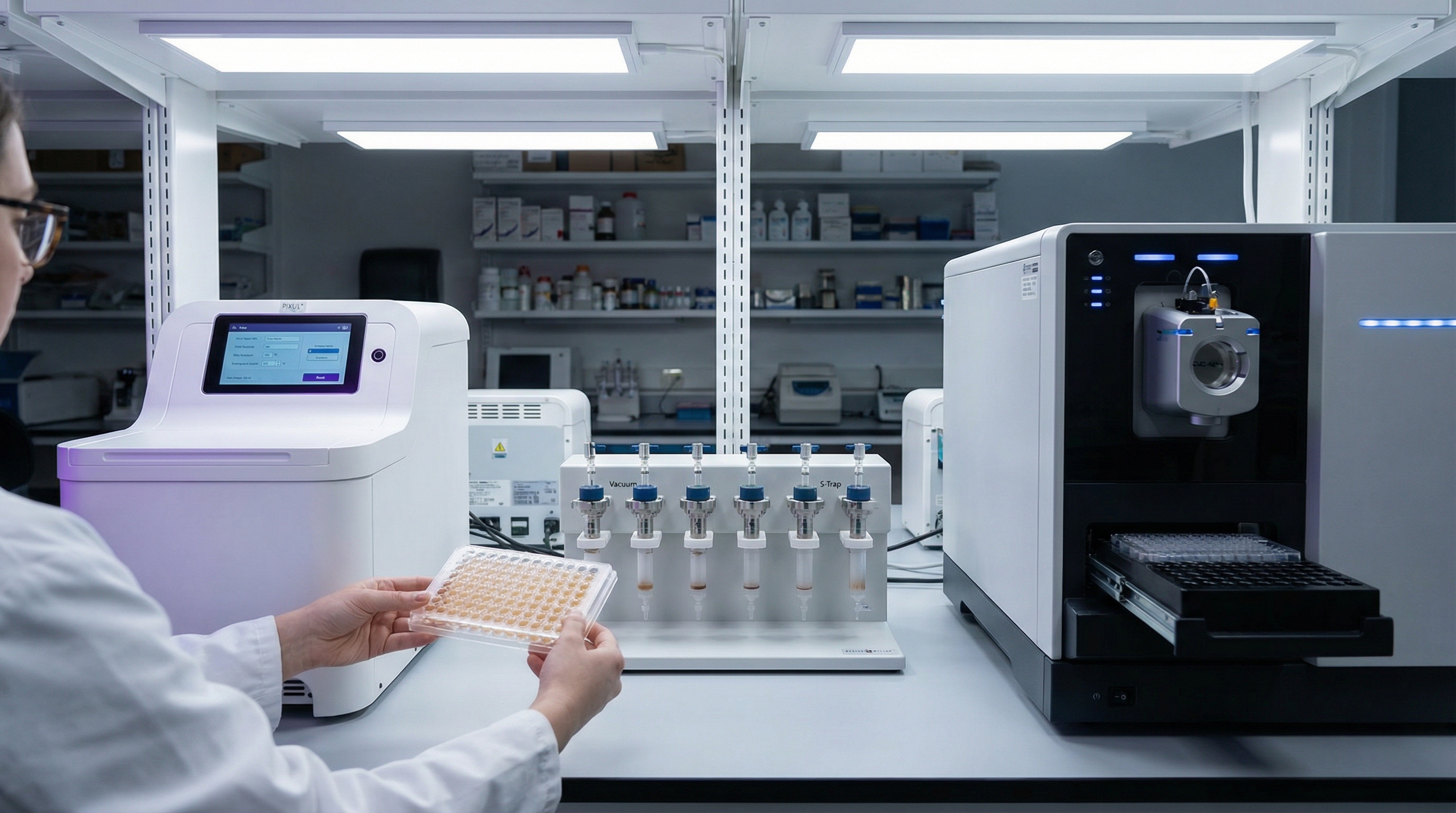

Current approaches to proteomics sample prep
Each method has its place. Each breaks at scale.
75% of proteomics error originates from sample processing. 72% from extraction alone.
Probe sonicators
The lab default. One sample at a time, operator-dependent results, localized thermal spikes that degrade proteins, and cross-contamination risk between samples.
Bead beaters
Handle tissue disruption at scale, but introduce bead contamination into LC-MS/MS samples. Heat generation degrades proteins. Variable extraction quality between wells.
Focused ultrasonicators
Deliver precision but at ~$400 per proprietary plate and sequential processing that takes 7+ hours for 96 samples. Vendor lock-in compounds the operating cost.
Chemical lysis
Simple and inexpensive but incomplete, with particularly poor recovery of membrane-bound and nuclear proteins. Published data confirms sonication significantly improves detection of these protein classes.
Full liquid handling automation
Solves throughput but costs $150K–$300K and requires dedicated lab space, specialized operators, and extensive validation.
What if your sample prep matched your mass spec throughput?
The ideal solution would process 96 samples in parallel, deliver reproducible protein extraction validated by LC-MS/MS, handle any sample type from FFPE to fresh cells, and use standard consumables at $2 per plate.

PIXUL: High-throughput sonicator for proteomics sample preparation
PIXUL processes 96 samples in parallel, in 10–30 minutes, in a standard $2 plate. Combined with S-Trap or SP3 digestion, it is the fastest path from cells to peptides, with the reproducibility your data demands.
Built by researchers at the University of Washington. Licensed and supported by Active Motif.
Your mass spec is fast. Your sample prep should be too.
PIXUL processes 96 samples in parallel, so your sample preparation finally matches your instrument throughput. Load samples in a standard microplate, set conditions on the touchscreen, and walk away. Combined with S-Trap or SP3 digestion, it is the fastest cells-to-peptides workflow available.
Sample prep causes 75% of proteomics error. PIXUL cuts it at the source.
Published research shows that sample processing causes 75% of total error in proteomics, with 72% from extraction alone. PIXUL's arrayed 2 MHz transducers deliver uniform cavitation in every well, independently and simultaneously. No positional effects. No operator variation.
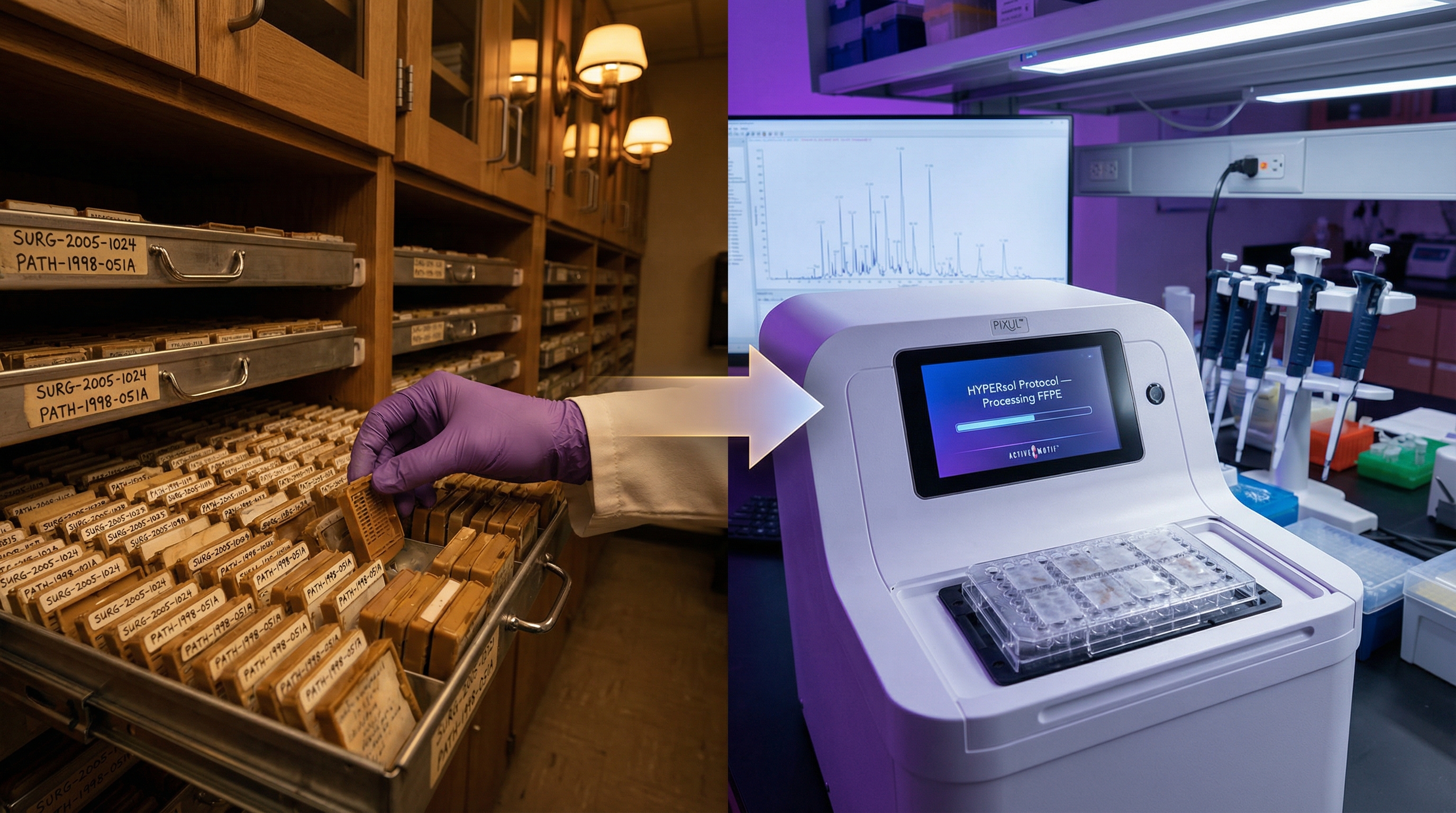
From FFPE archives to fresh cell lines, all in the same plate.
Load your FFPE, frozen, and cell samples on the same 96-well plate, assign up to 12 different sonication conditions by column, and process them all simultaneously. Combined with HYPERsol and S-Trap, PIXUL unlocks your tissue archives for high-throughput proteomics.
Protein identification: PIXUL vs. probe sonication
Head-to-head LC-MS/MS validation from PIXUL-prepared mouse liver tissues shows equivalent protein identification depth with maintained quality control metrics.
Equivalent depth. 96x the throughput.
Missed Cleavages
Within normal range
N-Terminal Acetylation
Within normal range
Oxidized Methionine
Within normal range
Source: Active Motif LC-MS/MS technical documentation. Mouse liver tissue samples.
Proteomics workflow with PIXUL
Standard plate format at every step. No proprietary consumables. No sample transfers between vessels.
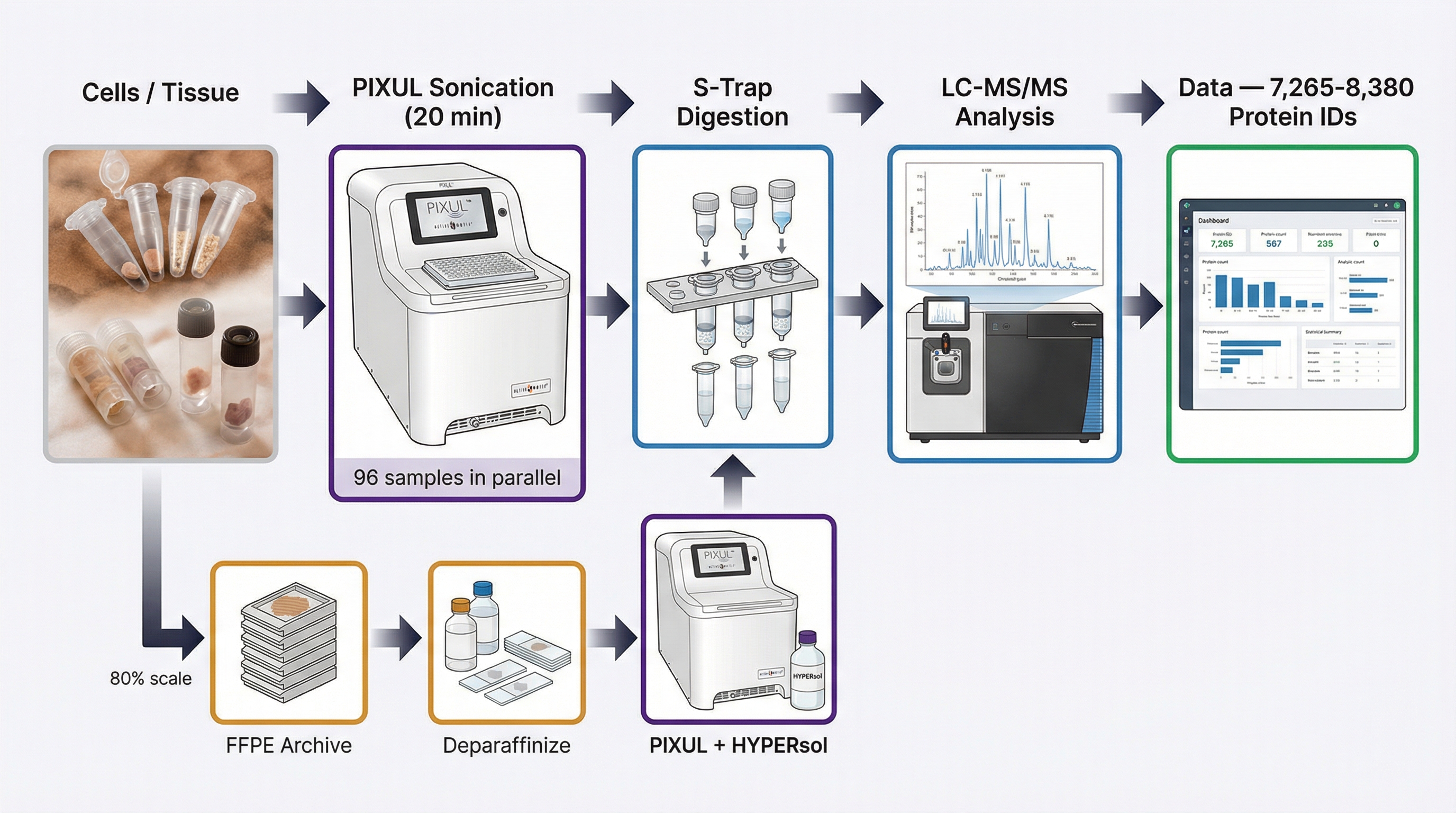
Standard workflow
FFPE workflow

PIXUL for proteomics
Compact, walk-away operation with no proprietary consumables.
| Throughput | 1–96 samples per run |
| Run Time | 10–30 minutes |
| Plate Format | Standard 96-well round-bottom |
| Frequency | 2 MHz megasonication |
| Conditions per Plate | Up to 12 |
| Dimensions | 45 × 59 × 34 cm |
| Weight | 23 kg |
| Setup Time | 15 minutes |
| Consumable Cost | ~$2/plate (standard) |

The data speaks for itself
Approximately one-third of PIXUL installations are proteomics labs, driven entirely by word-of-mouth with zero active marketing. Pharmaceutical companies are repeat purchasers.
Proteomics installs
Of all PIXUL installations, from word-of-mouth alone
Protein IDs (PIXUL)
LC-MS/MS validated, equivalent to probe sonication
Publications
Peer-reviewed validation across all applications
Per plate
Standard consumables vs. ~$400 proprietary
Since the first time that I sonicated my samples and I got the expected results all in ~30 min, I don't want to look back at those days when I had to hold my tube and be seated for hours sonicating all my samples one by one.
Dr. Alexander Schmidt Webinar: PIXUL in Proteomics Workflows
Independent validation from Dr. Alexander Schmidt's laboratory, confirming PIXUL's performance in high-throughput proteomics sample preparation. The webinar covers integration with established LC-MS/MS pipelines and real-world throughput data from an active proteomics core facility.
Pharmaceutical companies are among the repeat purchasers of PIXUL, deploying multiple units across proteomics core facilities.
Works with your existing pipeline
PIXUL integrates directly with the proteomics workflows you already use. No protocol changes required.
S-Trap
ProtiFi partnership validated
Si-Trap
Compatible digestion workflow
SP3
Bead-based sample processing
HYPERsol
FFPE proteomics protocol
Ready to accelerate your proteomics workflow?
See how PIXUL fits into your LC-MS/MS pipeline. Test with your own samples on your own mass spec platform.